W3cubDocs
/scikit-learnsklearn.ensemble.RandomTreesEmbedding
-
class sklearn.ensemble.RandomTreesEmbedding(n_estimators=10, max_depth=5, min_samples_split=2, min_samples_leaf=1, min_weight_fraction_leaf=0.0, max_leaf_nodes=None, min_impurity_split=1e-07, sparse_output=True, n_jobs=1, random_state=None, verbose=0, warm_start=False)[source] -
An ensemble of totally random trees.
An unsupervised transformation of a dataset to a high-dimensional sparse representation. A datapoint is coded according to which leaf of each tree it is sorted into. Using a one-hot encoding of the leaves, this leads to a binary coding with as many ones as there are trees in the forest.
The dimensionality of the resulting representation is
n_out <= n_estimators * max_leaf_nodes. Ifmax_leaf_nodes == None, the number of leaf nodes is at mostn_estimators * 2 ** max_depth.Read more in the User Guide.
Parameters: n_estimators : integer, optional (default=10)
Number of trees in the forest.
max_depth : integer, optional (default=5)
The maximum depth of each tree. If None, then nodes are expanded until all leaves are pure or until all leaves contain less than min_samples_split samples.
min_samples_split : int, float, optional (default=2)
The minimum number of samples required to split an internal node:
- If int, then consider
min_samples_splitas the minimum number. - If float, then
min_samples_splitis a percentage andceil(min_samples_split * n_samples)is the minimum number of samples for each split.
Changed in version 0.18: Added float values for percentages.
min_samples_leaf : int, float, optional (default=1)
The minimum number of samples required to be at a leaf node:
- If int, then consider
min_samples_leafas the minimum number. - If float, then
min_samples_leafis a percentage andceil(min_samples_leaf * n_samples)is the minimum number of samples for each node.
Changed in version 0.18: Added float values for percentages.
min_weight_fraction_leaf : float, optional (default=0.)
The minimum weighted fraction of the input samples required to be at a leaf node.
max_leaf_nodes : int or None, optional (default=None)
Grow trees with
max_leaf_nodesin best-first fashion. Best nodes are defined as relative reduction in impurity. If None then unlimited number of leaf nodes.min_impurity_split : float, optional (default=1e-7)
Threshold for early stopping in tree growth. A node will split if its impurity is above the threshold, otherwise it is a leaf.
New in version 0.18.
sparse_output : bool, optional (default=True)
Whether or not to return a sparse CSR matrix, as default behavior, or to return a dense array compatible with dense pipeline operators.
n_jobs : integer, optional (default=1)
The number of jobs to run in parallel for both
fitandpredict. If -1, then the number of jobs is set to the number of cores.random_state : int, RandomState instance or None, optional (default=None)
If int, random_state is the seed used by the random number generator; If RandomState instance, random_state is the random number generator; If None, the random number generator is the RandomState instance used by
np.random.verbose : int, optional (default=0)
Controls the verbosity of the tree building process.
warm_start : bool, optional (default=False)
When set to
True, reuse the solution of the previous call to fit and add more estimators to the ensemble, otherwise, just fit a whole new forest.Attributes: estimators_ : list of DecisionTreeClassifier
The collection of fitted sub-estimators.
References
[R165] P. Geurts, D. Ernst., and L. Wehenkel, “Extremely randomized trees”, Machine Learning, 63(1), 3-42, 2006. [R166] Moosmann, F. and Triggs, B. and Jurie, F. “Fast discriminative visual codebooks using randomized clustering forests” NIPS 2007 Methods
apply(X)Apply trees in the forest to X, return leaf indices. decision_path(X)Return the decision path in the forest fit(X[, y, sample_weight])Fit estimator. fit_transform(X[, y, sample_weight])Fit estimator and transform dataset. get_params([deep])Get parameters for this estimator. set_params(**params)Set the parameters of this estimator. transform(X)Transform dataset. -
__init__(n_estimators=10, max_depth=5, min_samples_split=2, min_samples_leaf=1, min_weight_fraction_leaf=0.0, max_leaf_nodes=None, min_impurity_split=1e-07, sparse_output=True, n_jobs=1, random_state=None, verbose=0, warm_start=False)[source]
-
apply(X)[source] -
Apply trees in the forest to X, return leaf indices.
Parameters: X : array-like or sparse matrix, shape = [n_samples, n_features]
The input samples. Internally, its dtype will be converted to
dtype=np.float32. If a sparse matrix is provided, it will be converted into a sparsecsr_matrix.Returns: X_leaves : array_like, shape = [n_samples, n_estimators]
For each datapoint x in X and for each tree in the forest, return the index of the leaf x ends up in.
-
decision_path(X)[source] -
Return the decision path in the forest
New in version 0.18.
Parameters: X : array-like or sparse matrix, shape = [n_samples, n_features]
The input samples. Internally, its dtype will be converted to
dtype=np.float32. If a sparse matrix is provided, it will be converted into a sparsecsr_matrix.Returns: indicator : sparse csr array, shape = [n_samples, n_nodes]
Return a node indicator matrix where non zero elements indicates that the samples goes through the nodes.
n_nodes_ptr : array of size (n_estimators + 1, )
The columns from indicator[n_nodes_ptr[i]:n_nodes_ptr[i+1]] gives the indicator value for the i-th estimator.
-
feature_importances_ -
- Return the feature importances (the higher, the more important the
- feature).
Returns: feature_importances_ : array, shape = [n_features]
-
fit(X, y=None, sample_weight=None)[source] -
Fit estimator.
Parameters: X : array-like or sparse matrix, shape=(n_samples, n_features)
The input samples. Use
dtype=np.float32for maximum efficiency. Sparse matrices are also supported, use sparsecsc_matrixfor maximum efficiency.Returns: self : object
Returns self.
-
fit_transform(X, y=None, sample_weight=None)[source] -
Fit estimator and transform dataset.
Parameters: X : array-like or sparse matrix, shape=(n_samples, n_features)
Input data used to build forests. Use
dtype=np.float32for maximum efficiency.Returns: X_transformed : sparse matrix, shape=(n_samples, n_out)
Transformed dataset.
-
get_params(deep=True)[source] -
Get parameters for this estimator.
Parameters: deep: boolean, optional :
If True, will return the parameters for this estimator and contained subobjects that are estimators.
Returns: params : mapping of string to any
Parameter names mapped to their values.
-
set_params(**params)[source] -
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as pipelines). The latter have parameters of the form
<component>__<parameter>so that it’s possible to update each component of a nested object.Returns: self :
-
transform(X)[source] -
Transform dataset.
Parameters: X : array-like or sparse matrix, shape=(n_samples, n_features)
Input data to be transformed. Use
dtype=np.float32for maximum efficiency. Sparse matrices are also supported, use sparsecsr_matrixfor maximum efficiency.Returns: X_transformed : sparse matrix, shape=(n_samples, n_out)
Transformed dataset.
- If int, then consider
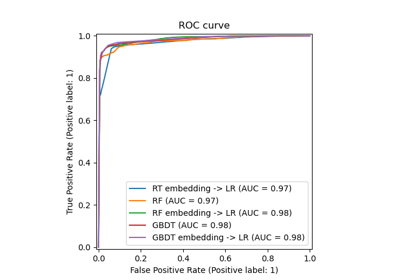
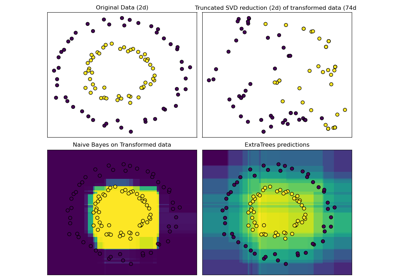
Examples using sklearn.ensemble.RandomTreesEmbedding
© 2007–2016 The scikit-learn developers
Licensed under the 3-clause BSD License.
http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomTreesEmbedding.html