W3cubDocs
/scikit-learnMulti-dimensional scaling
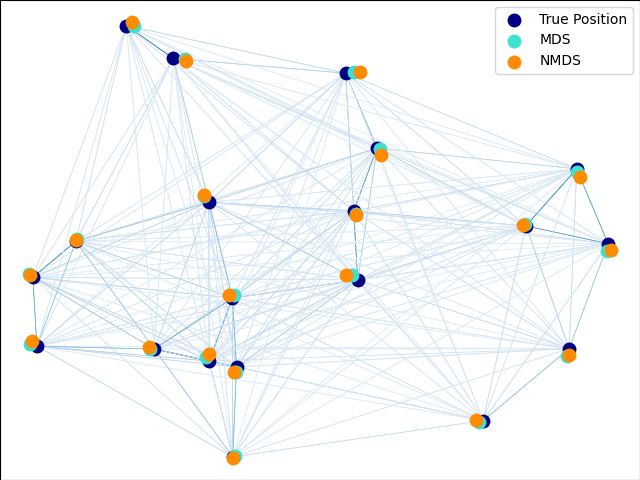
An illustration of the metric and non-metric MDS on generated noisy data.
The reconstructed points using the metric MDS and non metric MDS are slightly shifted to avoid overlapping.
# Author: Nelle Varoquaux <[email protected]> # License: BSD print(__doc__) import numpy as np from matplotlib import pyplot as plt from matplotlib.collections import LineCollection from sklearn import manifold from sklearn.metrics import euclidean_distances from sklearn.decomposition import PCA n_samples = 20 seed = np.random.RandomState(seed=3) X_true = seed.randint(0, 20, 2 * n_samples).astype(np.float) X_true = X_true.reshape((n_samples, 2)) # Center the data X_true -= X_true.mean() similarities = euclidean_distances(X_true) # Add noise to the similarities noise = np.random.rand(n_samples, n_samples) noise = noise + noise.T noise[np.arange(noise.shape[0]), np.arange(noise.shape[0])] = 0 similarities += noise mds = manifold.MDS(n_components=2, max_iter=3000, eps=1e-9, random_state=seed, dissimilarity="precomputed", n_jobs=1) pos = mds.fit(similarities).embedding_ nmds = manifold.MDS(n_components=2, metric=False, max_iter=3000, eps=1e-12, dissimilarity="precomputed", random_state=seed, n_jobs=1, n_init=1) npos = nmds.fit_transform(similarities, init=pos) # Rescale the data pos *= np.sqrt((X_true ** 2).sum()) / np.sqrt((pos ** 2).sum()) npos *= np.sqrt((X_true ** 2).sum()) / np.sqrt((npos ** 2).sum()) # Rotate the data clf = PCA(n_components=2) X_true = clf.fit_transform(X_true) pos = clf.fit_transform(pos) npos = clf.fit_transform(npos) fig = plt.figure(1) ax = plt.axes([0., 0., 1., 1.]) s = 100 plt.scatter(X_true[:, 0], X_true[:, 1], color='navy', s=s, lw=0, label='True Position') plt.scatter(pos[:, 0], pos[:, 1], color='turquoise', s=s, lw=0, label='MDS') plt.scatter(npos[:, 0], npos[:, 1], color='darkorange', s=s, lw=0, label='NMDS') plt.legend(scatterpoints=1, loc='best', shadow=False) similarities = similarities.max() / similarities * 100 similarities[np.isinf(similarities)] = 0 # Plot the edges start_idx, end_idx = np.where(pos) # a sequence of (*line0*, *line1*, *line2*), where:: # linen = (x0, y0), (x1, y1), ... (xm, ym) segments = [[X_true[i, :], X_true[j, :]] for i in range(len(pos)) for j in range(len(pos))] values = np.abs(similarities) lc = LineCollection(segments, zorder=0, cmap=plt.cm.Blues, norm=plt.Normalize(0, values.max())) lc.set_array(similarities.flatten()) lc.set_linewidths(0.5 * np.ones(len(segments))) ax.add_collection(lc) plt.show()
Total running time of the script: (0 minutes 0.183 seconds)
Download Python source code:
plot_mds.py
Download IPython notebook:
plot_mds.ipynb
© 2007–2016 The scikit-learn developers
Licensed under the 3-clause BSD License.
http://scikit-learn.org/stable/auto_examples/manifold/plot_mds.html