W3cubDocs
/scikit-learnModel Complexity Influence
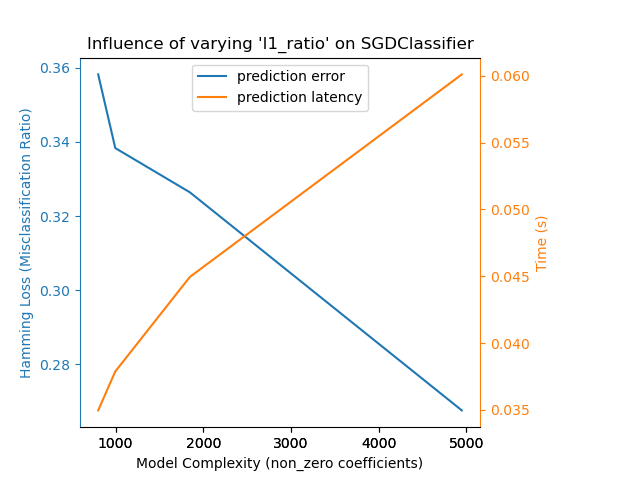
Demonstrate how model complexity influences both prediction accuracy and computational performance.
The dataset is the Boston Housing dataset (resp. 20 Newsgroups) for regression (resp. classification).
For each class of models we make the model complexity vary through the choice of relevant model parameters and measure the influence on both computational performance (latency) and predictive power (MSE or Hamming Loss).
print(__doc__) # Author: Eustache Diemert <[email protected]> # License: BSD 3 clause import time import numpy as np import matplotlib.pyplot as plt from mpl_toolkits.axes_grid1.parasite_axes import host_subplot from mpl_toolkits.axisartist.axislines import Axes from scipy.sparse.csr import csr_matrix from sklearn import datasets from sklearn.utils import shuffle from sklearn.metrics import mean_squared_error from sklearn.svm.classes import NuSVR from sklearn.ensemble.gradient_boosting import GradientBoostingRegressor from sklearn.linear_model.stochastic_gradient import SGDClassifier from sklearn.metrics import hamming_loss
Routines
# initialize random generator
np.random.seed(0)
def generate_data(case, sparse=False):
"""Generate regression/classification data."""
bunch = None
if case == 'regression':
bunch = datasets.load_boston()
elif case == 'classification':
bunch = datasets.fetch_20newsgroups_vectorized(subset='all')
X, y = shuffle(bunch.data, bunch.target)
offset = int(X.shape[0] * 0.8)
X_train, y_train = X[:offset], y[:offset]
X_test, y_test = X[offset:], y[offset:]
if sparse:
X_train = csr_matrix(X_train)
X_test = csr_matrix(X_test)
else:
X_train = np.array(X_train)
X_test = np.array(X_test)
y_test = np.array(y_test)
y_train = np.array(y_train)
data = {'X_train': X_train, 'X_test': X_test, 'y_train': y_train,
'y_test': y_test}
return data
def benchmark_influence(conf):
"""
Benchmark influence of :changing_param: on both MSE and latency.
"""
prediction_times = []
prediction_powers = []
complexities = []
for param_value in conf['changing_param_values']:
conf['tuned_params'][conf['changing_param']] = param_value
estimator = conf['estimator'](**conf['tuned_params'])
print("Benchmarking %s" % estimator)
estimator.fit(conf['data']['X_train'], conf['data']['y_train'])
conf['postfit_hook'](estimator)
complexity = conf['complexity_computer'](estimator)
complexities.append(complexity)
start_time = time.time()
for _ in range(conf['n_samples']):
y_pred = estimator.predict(conf['data']['X_test'])
elapsed_time = (time.time() - start_time) / float(conf['n_samples'])
prediction_times.append(elapsed_time)
pred_score = conf['prediction_performance_computer'](
conf['data']['y_test'], y_pred)
prediction_powers.append(pred_score)
print("Complexity: %d | %s: %.4f | Pred. Time: %fs\n" % (
complexity, conf['prediction_performance_label'], pred_score,
elapsed_time))
return prediction_powers, prediction_times, complexities
def plot_influence(conf, mse_values, prediction_times, complexities):
"""
Plot influence of model complexity on both accuracy and latency.
"""
plt.figure(figsize=(12, 6))
host = host_subplot(111, axes_class=Axes)
plt.subplots_adjust(right=0.75)
par1 = host.twinx()
host.set_xlabel('Model Complexity (%s)' % conf['complexity_label'])
y1_label = conf['prediction_performance_label']
y2_label = "Time (s)"
host.set_ylabel(y1_label)
par1.set_ylabel(y2_label)
p1, = host.plot(complexities, mse_values, 'b-', label="prediction error")
p2, = par1.plot(complexities, prediction_times, 'r-',
label="latency")
host.legend(loc='upper right')
host.axis["left"].label.set_color(p1.get_color())
par1.axis["right"].label.set_color(p2.get_color())
plt.title('Influence of Model Complexity - %s' % conf['estimator'].__name__)
plt.show()
def _count_nonzero_coefficients(estimator):
a = estimator.coef_.toarray()
return np.count_nonzero(a)
main code
regression_data = generate_data('regression')
classification_data = generate_data('classification', sparse=True)
configurations = [
{'estimator': SGDClassifier,
'tuned_params': {'penalty': 'elasticnet', 'alpha': 0.001, 'loss':
'modified_huber', 'fit_intercept': True},
'changing_param': 'l1_ratio',
'changing_param_values': [0.25, 0.5, 0.75, 0.9],
'complexity_label': 'non_zero coefficients',
'complexity_computer': _count_nonzero_coefficients,
'prediction_performance_computer': hamming_loss,
'prediction_performance_label': 'Hamming Loss (Misclassification Ratio)',
'postfit_hook': lambda x: x.sparsify(),
'data': classification_data,
'n_samples': 30},
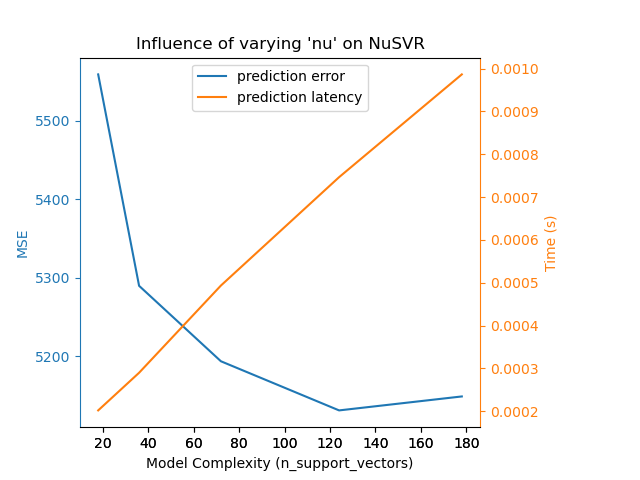
{'estimator': NuSVR,
'tuned_params': {'C': 1e3, 'gamma': 2 ** -15},
'changing_param': 'nu',
'changing_param_values': [0.1, 0.25, 0.5, 0.75, 0.9],
'complexity_label': 'n_support_vectors',
'complexity_computer': lambda x: len(x.support_vectors_),
'data': regression_data,
'postfit_hook': lambda x: x,
'prediction_performance_computer': mean_squared_error,
'prediction_performance_label': 'MSE',
'n_samples': 30},
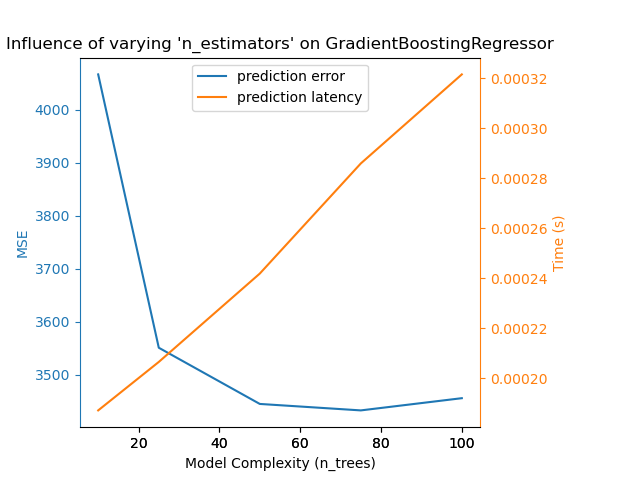
{'estimator': GradientBoostingRegressor,
'tuned_params': {'loss': 'ls'},
'changing_param': 'n_estimators',
'changing_param_values': [10, 50, 100, 200, 500],
'complexity_label': 'n_trees',
'complexity_computer': lambda x: x.n_estimators,
'data': regression_data,
'postfit_hook': lambda x: x,
'prediction_performance_computer': mean_squared_error,
'prediction_performance_label': 'MSE',
'n_samples': 30},
]
for conf in configurations:
prediction_performances, prediction_times, complexities = \
benchmark_influence(conf)
plot_influence(conf, prediction_performances, prediction_times,
complexities)
Out:
Benchmarking SGDClassifier(alpha=0.001, average=False, class_weight=None, epsilon=0.1,
eta0=0.0, fit_intercept=True, l1_ratio=0.25,
learning_rate='optimal', loss='modified_huber', n_iter=5, n_jobs=1,
penalty='elasticnet', power_t=0.5, random_state=None, shuffle=True,
verbose=0, warm_start=False)
Complexity: 4454 | Hamming Loss (Misclassification Ratio): 0.2501 | Pred. Time: 0.027734s
Benchmarking SGDClassifier(alpha=0.001, average=False, class_weight=None, epsilon=0.1,
eta0=0.0, fit_intercept=True, l1_ratio=0.5, learning_rate='optimal',
loss='modified_huber', n_iter=5, n_jobs=1, penalty='elasticnet',
power_t=0.5, random_state=None, shuffle=True, verbose=0,
warm_start=False)
Complexity: 1624 | Hamming Loss (Misclassification Ratio): 0.2923 | Pred. Time: 0.019370s
Benchmarking SGDClassifier(alpha=0.001, average=False, class_weight=None, epsilon=0.1,
eta0=0.0, fit_intercept=True, l1_ratio=0.75,
learning_rate='optimal', loss='modified_huber', n_iter=5, n_jobs=1,
penalty='elasticnet', power_t=0.5, random_state=None, shuffle=True,
verbose=0, warm_start=False)
Complexity: 873 | Hamming Loss (Misclassification Ratio): 0.3191 | Pred. Time: 0.015396s
Benchmarking SGDClassifier(alpha=0.001, average=False, class_weight=None, epsilon=0.1,
eta0=0.0, fit_intercept=True, l1_ratio=0.9, learning_rate='optimal',
loss='modified_huber', n_iter=5, n_jobs=1, penalty='elasticnet',
power_t=0.5, random_state=None, shuffle=True, verbose=0,
warm_start=False)
Complexity: 655 | Hamming Loss (Misclassification Ratio): 0.3252 | Pred. Time: 0.014888s
Benchmarking NuSVR(C=1000.0, cache_size=200, coef0=0.0, degree=3, gamma=3.0517578125e-05,
kernel='rbf', max_iter=-1, nu=0.1, shrinking=True, tol=0.001,
verbose=False)
Complexity: 69 | MSE: 31.8133 | Pred. Time: 0.000360s
Benchmarking NuSVR(C=1000.0, cache_size=200, coef0=0.0, degree=3, gamma=3.0517578125e-05,
kernel='rbf', max_iter=-1, nu=0.25, shrinking=True, tol=0.001,
verbose=False)
Complexity: 136 | MSE: 25.6140 | Pred. Time: 0.000645s
Benchmarking NuSVR(C=1000.0, cache_size=200, coef0=0.0, degree=3, gamma=3.0517578125e-05,
kernel='rbf', max_iter=-1, nu=0.5, shrinking=True, tol=0.001,
verbose=False)
Complexity: 243 | MSE: 22.3315 | Pred. Time: 0.001106s
Benchmarking NuSVR(C=1000.0, cache_size=200, coef0=0.0, degree=3, gamma=3.0517578125e-05,
kernel='rbf', max_iter=-1, nu=0.75, shrinking=True, tol=0.001,
verbose=False)
Complexity: 350 | MSE: 21.3679 | Pred. Time: 0.001699s
Benchmarking NuSVR(C=1000.0, cache_size=200, coef0=0.0, degree=3, gamma=3.0517578125e-05,
kernel='rbf', max_iter=-1, nu=0.9, shrinking=True, tol=0.001,
verbose=False)
Complexity: 404 | MSE: 21.0915 | Pred. Time: 0.001865s
Benchmarking GradientBoostingRegressor(alpha=0.9, criterion='friedman_mse', init=None,
learning_rate=0.1, loss='ls', max_depth=3, max_features=None,
max_leaf_nodes=None, min_impurity_split=1e-07,
min_samples_leaf=1, min_samples_split=2,
min_weight_fraction_leaf=0.0, n_estimators=10, presort='auto',
random_state=None, subsample=1.0, verbose=0, warm_start=False)
Complexity: 10 | MSE: 28.9793 | Pred. Time: 0.000111s
Benchmarking GradientBoostingRegressor(alpha=0.9, criterion='friedman_mse', init=None,
learning_rate=0.1, loss='ls', max_depth=3, max_features=None,
max_leaf_nodes=None, min_impurity_split=1e-07,
min_samples_leaf=1, min_samples_split=2,
min_weight_fraction_leaf=0.0, n_estimators=50, presort='auto',
random_state=None, subsample=1.0, verbose=0, warm_start=False)
Complexity: 50 | MSE: 8.3398 | Pred. Time: 0.000187s
Benchmarking GradientBoostingRegressor(alpha=0.9, criterion='friedman_mse', init=None,
learning_rate=0.1, loss='ls', max_depth=3, max_features=None,
max_leaf_nodes=None, min_impurity_split=1e-07,
min_samples_leaf=1, min_samples_split=2,
min_weight_fraction_leaf=0.0, n_estimators=100,
presort='auto', random_state=None, subsample=1.0, verbose=0,
warm_start=False)
Complexity: 100 | MSE: 7.0096 | Pred. Time: 0.000267s
Benchmarking GradientBoostingRegressor(alpha=0.9, criterion='friedman_mse', init=None,
learning_rate=0.1, loss='ls', max_depth=3, max_features=None,
max_leaf_nodes=None, min_impurity_split=1e-07,
min_samples_leaf=1, min_samples_split=2,
min_weight_fraction_leaf=0.0, n_estimators=200,
presort='auto', random_state=None, subsample=1.0, verbose=0,
warm_start=False)
Complexity: 200 | MSE: 6.1836 | Pred. Time: 0.000420s
Benchmarking GradientBoostingRegressor(alpha=0.9, criterion='friedman_mse', init=None,
learning_rate=0.1, loss='ls', max_depth=3, max_features=None,
max_leaf_nodes=None, min_impurity_split=1e-07,
min_samples_leaf=1, min_samples_split=2,
min_weight_fraction_leaf=0.0, n_estimators=500,
presort='auto', random_state=None, subsample=1.0, verbose=0,
warm_start=False)
Complexity: 500 | MSE: 6.3426 | Pred. Time: 0.000911s
Total running time of the script: (0 minutes 25.008 seconds)
Download Python source code:
plot_model_complexity_influence.py
Download IPython notebook:
plot_model_complexity_influence.ipynb
© 2007–2016 The scikit-learn developers
Licensed under the 3-clause BSD License.
http://scikit-learn.org/stable/auto_examples/applications/plot_model_complexity_influence.html